Introduction
Cytogenetics was introduced in 1900 as a tool to determine the correct number of chromosomes of an organism. Since the 1970s, it has been used to detect clinical syndromes; it helped to understand the chromosomal aberrations and phenotypes associated with it.
Karyotyping was the first cytogenetics tool used to study the characteristics of chromosomes. But, the resolution of this method was not enough to study the precise position and function of a gene in a chromosome. The limitations[7] of karyotyping were: (1) it could only analyze >3 MB of DNA, (2) the process required mitotically active cells, (3) it was difficult to study the complex chromosomal rearrangements, and (4) preparation of slides were required (which made it a time taking process).
Later in the 1980s, molecular cytogenetic techniques were introduced, which brought a different era of analyzing chromosomes with variant colors. This was the first technique to introduce the concept of “in situ hybridization”. It helped the researchers in different ways[7]: (1) by boosting the chromosome mapping strategies, (2) permitted the identification of complex chromosome rearrangements, and (3) helped in the identification of chromosomal aberrations such as translocation, deletion, insertion, and inversion.
Techniques in Molecular Cytogenetics
Fluorescent in situ Hybridization (FISH)
Pinkel introduced the Fluorescent in situ hybridization technique in 1986; he used fluorescently labeled probes to analyze the genomic DNA/RNA region for any abnormalities. The labeled probes bind and glow to their complementary regions on the chromosome – this can be observed using a fluorescent microscope. Molecular cytogenetics has paved the way for clinical and research developments by introducing the FISH technique to diagnose and treat abnormalities.
This method is highly efficient over the classic Karyotype technique, as it requires less time (it does not require cell culture) and provides a high-resolution view of the chromosomes. This technique allowed scientists to have a closer look at chromosomal aberrations and to diagnose the abnormalities with more accuracy and precision[7].
Method
DNA probes and target sequences are the two basic elements of FISH.
Probes
The choice of probes depends on the chromosomal regions to analyze. The three main types[7] of probes include:
- Whole-chromosome painting probes
- Derived from a single type of chromosome.
- By using degenerate oligonucleotide PCR, the probes are microdissected, amplified, and labeled to give a color to the whole chromosome.
- Mostly used to visualize the number of chromosomes at metaphase.
- It is not for the analysis of chromosomal aberrations.
- Repetitive sequence probes
- It highlights repetitive short sequences of chromosomal regions.
- Suitable to analyze the numerical aberrations in cells, either at metaphase or interphase.
- An example of these probes includes chromosome-specific centromeric and pan-telomeric probes.
- Pan-telomeric probes target the repeated (TTAGGG) sequences (present at the end of human chromosomes).
- Locus-specific (also known as a unique sequence) probes
- Target specific chromosomal sequences (present in only one copy of the chromosomes).
- Cloned into vectors for use, such as plasmids (1-10 kb), PAC, YAC, and BAC vectors.
- Used to analyze chromosome aberrations (in cells either at metaphase or interphase) such as deletion, inversion, and translocation.
Mechanism
- Prepare a probe (complementary to the sequence to be analyzed)
- Label the probes (either by nick translation, random priming, or PCR)
- Denature the labeled probe and target DNA
- Incubate probes with the target sequence of DNA to hybridize
- Analyze the probe binding regions using a fluorescent microscope
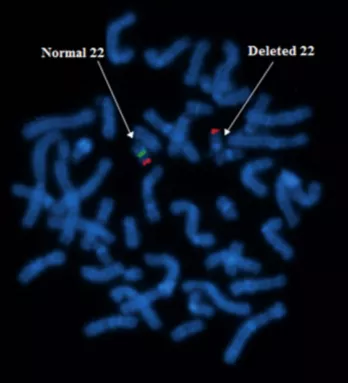
Figure: The image shows the deletion of chromosome 22 by using the FISH technique. Notice the absence of green color in the chromosome pointed by the arrow on the right-hand side.
Advantages
- Analyze large deletions and duplication in the germline.
- Diagnose Cancer (analyze the deletion, duplication, and translocation in the chromosomes)[3].
- Help in understanding whether cancer will respond to particular therapeutic drugs or not.
- Analyze mosaicism[3].
- One analysis can include several cells.
Limitations
- Adequate size of tissue is required.
- Pre-knowledge of the target is required.
- Changes in the copy number outside the target will not be identified[3].
The use of probes (in the FISH techniques) to analyze the chromosomes, requires the pre-knowledge of chromosomal aberration, which makes the analysis a complex process. To overcome this limitation, the FISH technique was modified several times for different diagnostic and research purposes based on the type of specimen to be analyzed or the information needed. Thus, genome-wide screening techniques were introduced, which produced precise and more reliable results compared to the previous techniques. These techniques involve:
- multicolor FISH (M-FISH, SKY, CCK)
- Primed in situ Labelling (PRINS)
- Comparative Genomic Hybridization (CGH)
The following sections will give a brief description of the modified FISH techniques with their limitations and advantages.
Comparative Genomic Hybridization (CGH)
CGH technique helps in the detection of copy number variation in the chromosome of an organism without the need for cultured cells. This technique (in comparison to FISH) is less time-consuming and provides an overview of specific chromosomal regions that have undergone deletion or duplication (which causes pathogenesis). It is more efficient in the analysis of oncogenes/cancer.
Method
CGH involves the labeling of DNA of interest and control samples with two different fluorescent probes. Like, label the test sample (tumor DNA) with green color and the control (normal sample) in the red color. Then, hybridize the metaphase chromosomes with both probes on the slide. The computer algorithm then gives the ratio of fluorophore generated by photometric analysis of the two DNA samples (it measures the ratio of test over the control or reference). The ratio provides an idea of deletion (ratio <1) or duplication (ratio >1) at specific chromosomal loci in examined DNA[11].
Array-CGH is the modified CGH technique. It is a chip-based technology that is less than an inch in size. Thousands of probes are present in a grid on the chip; differently colored test and control samples are mixed in the grid where they compete to bind with the available probes. The procedure and analysis to be done are the same as CGH. But, this technique (array-CGH) is more efficient than CGH, as it can simultaneously analyze large samples and multiple loci of the chromosomes in very less time.
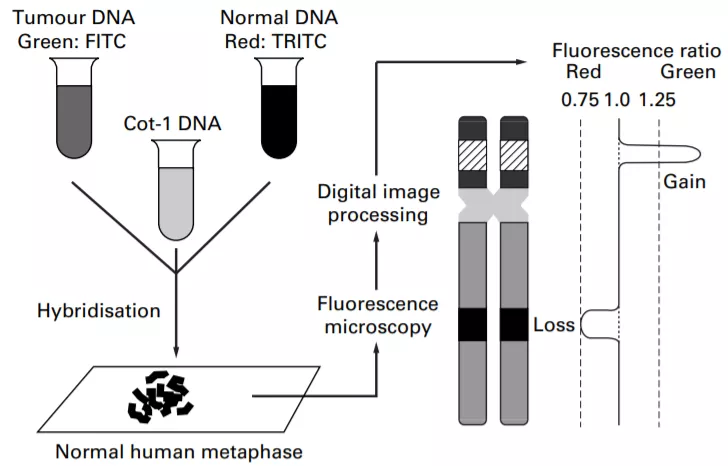
Figure 2: The figure shows the difference between the comparative genomic hybridization (CGH) and array-CGH. Notice how both determine the ratio of fluorophores and predict the gain or loss of a specific chromosomal locus.
Advantages
- Useful in cases where the mitotic index is low or null.
- It allows analyzing multiple genes simultaneously for the identification of copy number variants[3].
- Very efficient to analyze aneuploidy or polyploidy.
Limitations
- Difficult to analyze the balanced translocation/rearrangements and mosaicism[3].
- The resolution is only 10-20 Mb[2].
- Mutation in cells less than 20-30% can be missed by this approach[3].
Primed in situ Labelling (PRINS)
The primedin situ hybridization technique was first used to visualize the difference between the ɑ-satellite region of the two different chromosomes of humans. This technique is more sensitive and faster compared to all the other available FISH techniques[4]. A major difference between both techniques is; that FISH involves the use of labeled probes while PRINS involves the use of unlabeled probes.
Method
This technique involves the use of unlabeled probes. When the unlabeled probes are mixed with the test sample, labeled dNTPs, and DNA polymerase, probes bind with the specific complementary target sequence. After binding with the target sequence, they act as a primer, and a chain elongation reaction is catalyzed by the DNA polymerase (it uses labeled dNTPs) at hybridizing site. The reaction is catalyzed in three primary steps: (1) denaturation, (2) annealing, and (3) elongation. So, only the probe binding at the target site will be labeled.
The use of unlabeled probes in the PRINS technique helps in reducing the background disturbances/signals very efficiently. As only the probe binding to its target sequence will be polymerized and incorporate labeled dNTPs (which will give color to that particular chromosomal region)[4]. Moreover, this technique requires only 5-30 minutes for the reaction to occur. Because of the lesser time of the reaction, the morphology of the chromosome is well preserved.
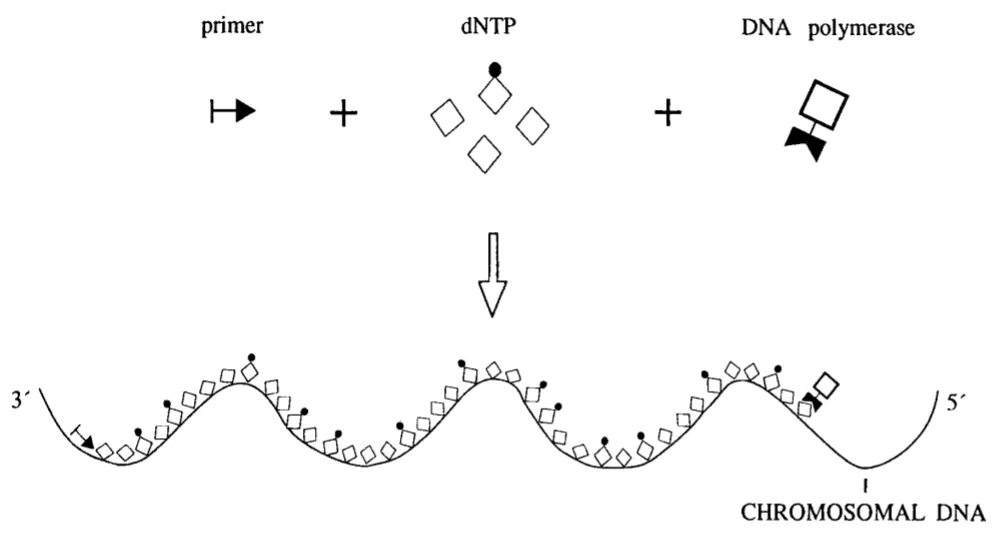
Figure 3: The image shows the PRINS reaction. The probe which binds at the target sequence act as primer and polymerase chain reaction only occurs at that place.
Advantages
- Fast process (requires only 5-30 minutes)
- Reduced background signals during the reaction[4].
- A large number of probes can be used for the analysis.
- Preserves chromosomal morphology[4].
Limitations
- Difficulty in identifying the targets that have low DNA or RNA copies.
Spectral Karyotyping(SKY) and Multiplex-Fluorescent in situ Hybridization (M-FISH) Technique
Spectral karyotyping (SKY) and M-FISH techniques are very advanced FISH-modified techniques. SKY technique was first introduced by Schröck E et al. in 1996. Both techniques involve the use of multicolor probes; 3-7 differently colored probes are used for single hybridization[6]. The only difference in both techniques is: SKY colors one entire chromosome in only one color whereas M-FISH highlights the regions of a chromosome in a different colored banding pattern. All 24 color combinations (generated from 3-5 differently colored probes) can be observed in the M-FISH technique. Presently, ‘SKY’ is being used to analyze the human, rat, and mouse chromosomes.
The procedures of both techniques are the same except for the software used to analyze the obtained images. That is why, in this section, only one technique – the SKY technique – will be discussed.
Method
Basic steps[10] involved in the SKY technique are given below:
- Make a cocktail of fluorescent-labeled probes with five fluorochromes such as FITC, Rhodamine, Texas Red, Cy5, and Cy5.5.
- Hybridize the metaphase chromosome with labeled probes and stain with 4,6-diamidino-2 phenylindole (DAPI) in an antifade medium.
- Use the spectral imaging system (such as The SpectraCube® Imaging system) to differentiate between the different spectral characteristics of chromosomes – chromosome-specific labeled probes (in different combinations) generate emission spectra of a specific chromosome, which are measured by the system.
- Use software (such as SKYView™ software) to analyze the spectra obtained from different fluorochrome combinations.
- The chromosomes are then arranged in a karyotype. To easily visualize the chromosomal aberrations, the software assigns a pseudocolor to the chromosomes.
- Capture the DAPI image separately and invert it into a G-banding pattern. It acts as a complement to spectral karyotyping, as it provides the banding pattern information.
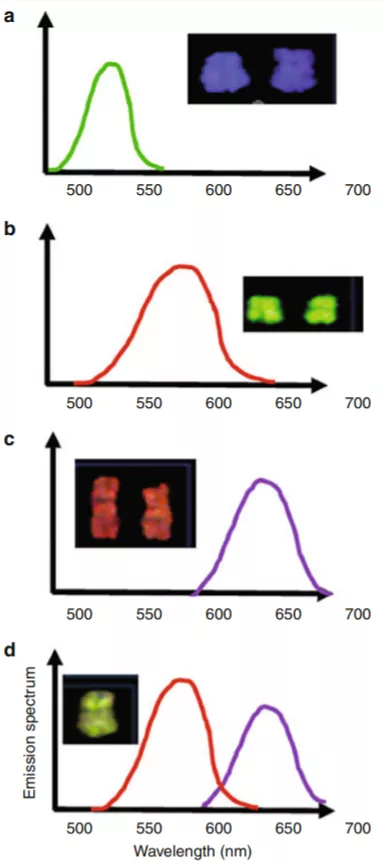
Figure 4: The image on the left-hand side shows how the software analyzes the spectral characteristic to identify chromosomes.
The image on the right-hand side shows DAP I (black and white[A]) stained and fluorochrome stained images [B]. [C] shows the karyotype of chromosomes stained in both (DAPI and fluorochrome) ways.
Advantages
- Used to analyze the interstitial deletions and cryptic translocations.
- Used in cancer cytogenetics, as it provides a detailed description of the abnormal karyotype and precisely defines the markers[6].
- It can be used to analyze any genome.
- Fast method.
Limitations
- It is difficult to detect the chromosomal rearrangements such as paracentric and pericentric inversion, small deletions, and duplications[6].
- The resolution is 1-3 Mb.
Application of Molecular Cytogenetics
-
FISH
- Breast Carcinomas: HER2 gene is the major cause of breast cancers in 10%-20% of cases. The FISH technique is used to analyze the overexpression of the HER2 gene[9].
- Pulmonary adenocarcinomas: In this case, rearrangement of Anaplastic lymphoma kinase (ALK) occurs. The rearrangement is due to the fusion of the EML4-ALK gene at chromosome 2p23. This fusion is observed by the FISH technique[9].
- Chronic Myeloid Leukemia (CML): This condition arises due to the Robertsonian translocation of the BCR/ABL1 genes. The FISH technique is used to detect this translocation and also to develop a targeted therapy for various carcinomas[9].
-
Comparative Genomic Hybridisation (CGH)
- Balanced structural rearrangements: Some chromosomal rearrangements (such as translocation between short arm of chromosome 11 and 12) are not easy to observe, as no phenotype difference can be seen due to the particular rearrangements, although, it can become lethal later. It can lead to a higher risk of cryptic genomic imbalances. So, array CGH is performed to visualize these translocations[8].
- Gain and loss of chromosomal regions: Deletions and duplications (copy number variants) can be easily observed by using the CGH technique. Example: in some cases, loss of chromosome 4 (q13.2-q21.1) and gain of chromosome 6 (q24.3) regions are observed using the CGH technique. Amplification of chromosomes 7, 16, and 19 are also studied using this method[8].
-
Primed in situ Labelling (PRINS)
- Telomeric rearrangements: By using the simple FISH technique, it is difficult to visualize the telomeric or subtelomeric deletions. For this purpose, the use of the PRINS technique is preferred, which produces the result in less than 3 hours.
- To visualize single-copy genes: single copy genes such as the SRY gene and SOX3 gene can be easily visualized using the PRINS technique.
-
Spectral Karyotyping (SKY) Technique
- Trisomy 11/22: A child with the trisomy of 11/22 can be detected by using the SKY technique, without examining its parents. A chromosome from an unknown origin is easy to get detected from this technique (different chromosomes colored with different fluorochromes)[1].
- 13q-Syndrome: Deletion at chromosome 13 can be observed by using the SKY technique but the point of breakage is difficult to analyze. To detect the breakage point, SKY with banding techniques is performed[1].
- Cancer: SKY has been used to diagnose various types of cancers, such as breast cancer, sarcomas, and carcinomas; to study the chromosomal aberration (cause of disease). The result of the analysis obtained from the SKY technique also helps to develop targeted therapy for the patients[1].
Conclusion
Molecular cytogenetics involve techniques based on the concept of in situ hybridization, such as Fluorescent in situ hybridization (FISH), spectral karyotyping (SKY), primed in situ hybridizations (PRINS), and comparative genomic hybridization (CGH). All these techniques involve the use of labeled probes, except PRINS in which labeled dNTPs (and unlabeled probes) are used instead of labeled probes. These techniques have made research easier and faster by providing reliable results in 1-3 hours and by allowing the analysis of multiple samples simultaneously. All these techniques have vast applications in clinical cytogenetics to diagnose various chromosomal aberrations (macro, as well as micro aberrations). Diagnosis and treatment (which help to develop targeted therapies) of cancer have also become easy since the evolution of these techniques. Molecular cytogenetics colored the world of chromosomes and it is expected that all the limitations will be overcome through the use of combinatorial approaches to these techniques.
References
- Bo Guo, Xiaoping Han, Zhanhe Wu, Wanming Da, Hongli Zhu (2014). Spectral karyotyping: a unique technique for the detection of complex genomic rearrangements in leukemia. Translational Pediatrics, 3(2), 135-139. DOI: 10.3978/j.issn.2224-4336.2014.01.02
- Gersen Steven L. and Keagle Martha B. (2013). The Principles of Clinical Cytogenetics (3rd ed.), Springer, New York.
- Gomes, A., & Korf, B. (2018). Genetic Testing Techniques. Pediatric Cancer Genetics, 47–64. DOI:10.1016/b978-0-323-48555-5.00005-3.
- Hindkjaer, J., Koch, J., Brandt, C., Kolvraa, S., & Bolund, L. (1996). Primed in situ labeling (PRINS). Molecular Biotechnology, 6(2), 201–211. DOI:10.1007/bf02740774.
- Hindkjaer, J., Bolund, L., & Kølvraa, S. (2001). Primed in Situ labeling. Cytometry: Part B, 55–68. DOI:10.1016/s0091-679x(01)64006-8
- Imataka, G., & Arisaka, O. (2011). Chromosome Analysis Using Spectral Karyotyping (SKY). Cell Biochemistry and Biophysics, 62(1), 13–17. DOI: 10.1007/s12013-011-9285-2.
- Kearney, L. (2001). Molecular cytogenetics. Best Practice & Research Clinical Haematology, 14(3), 645–668. DOI:10.1053/beha.2001.0159.
- Liu, J., Bernier, F., Lauzon, J., Lowry, R. B., & Chernos, J. (2011). Application of Microarray-Based Comparative Genomic Hybridization in Prenatal and Postnatal Settings: Three Case Reports. Genetics Research International, 1–9. DOI:10.4061/2011/976398
- Ratan, Z. A., Zaman, S. B., Mehta, V., Haidere, M. F., Runa, N. J., & Akter, N. (2017). Application of Fluorescence In Situ Hybridization (FISH) Technique for the Detection of Genetic Aberration in Medical Science. Cureus, 9(6). DOI:10.7759/cureus.1325
- Trakhtenbrot, L. (2011). Spectral Karyotyping. Encyclopedia of Cancer, 3472–3476. DOI:10.1007/978-3-642-16483-5_5433.
- Weiss M. M., Hermsen M. A., Meijer G. A., Grieken N. C. van, Baak J P., Kuipers E J, Diest P J van (1999). Comparative genomic hybridization. Journal of Clinical Pathology, 52, 243-251. DOI: 10.1136/mp.52.5.243.



