As an Amazon Associate Conductscience Inc earns revenue from qualifying purchases
Introduction
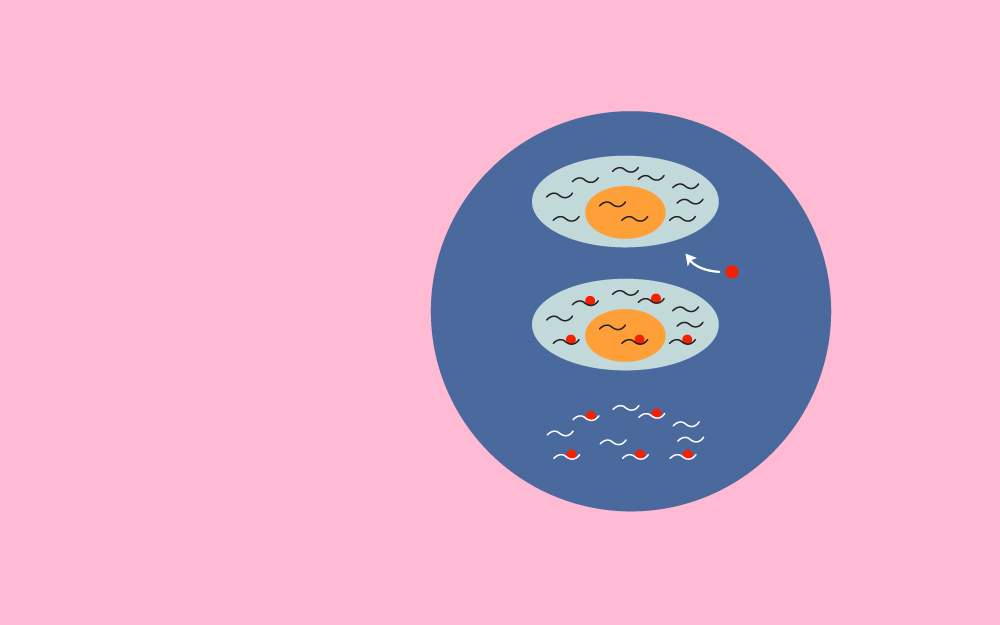
The pulse-chase analysis is a powerful technique to study the synthesis, processing, and transport of proteins. The pulse-chase analysis is widely used in biochemistry and molecular biology for the examination of the cellular process by exposing the cells to a labeled compound (pulse). The labeled compound is introduced into the molecule or pathway being studied. In the chase phase, an unlabeled form of that compound replaces the labeled compound. The reaction is monitored to note the time taken by the unlabeled form of the compound to replace the labeled form. These experiments allow the assessment and observation of the evolution of a biological process over time.
Principle
Pulse-chase analysis is a specialized form of metabolic labeling which employs radioactive amino acids for observing cellular events over time. The radioactive amino acids are added in the cells for a few minutes (the “pulse”), washed away, and then the cells are exposed to nonradioactive amino acids (in excess). The cells are then taken for protein extraction. Pulse-chase analysis is used for the study of protein synthesis and processing, the examination of intracellular localization of nascent proteins over time, and for the assessment of their secretion, translocation, and degradation.
Apparatus
The pulse-chase experiment consists of three stages: the actual pulse-chase, immunoprecipitation, and sodium-dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE). In the experiment, a fluorescently or radioactively tagged compound is used for time-based monitoring of the cellular events. After the pulse-chase, the proteins of interest are harvested by cell lysis through immunoprecipitation. The proteins are then run on SDS-PAGE for further analysis. The equipment needed for the experiment includes an incubator, Eppendorf tubes, a 26-gauge needle, a syringe, heat block, gel dryer, and Phosphor imaging screen.
Protocol (Magadán, 2017)
Radioactive pulse-chase
- Incubate the MDCK cells (grown and infected with 10 infectious doses of influenza A virus per cell for 5 hours at 37oC) with 5 ml Trypsin-EDTA at 37oC for 5 minutes.
- Transfer the cells to a 50 ml BD falcon tube and wash them twice with 10 ml pre-warmed Dulbecco’s phosphate-buffered saline (DPBS) and centrifuge for 1 minute at 2500 x g.
- Resuspend the cells in 1 ml DPBS and transfer in a 1.5 ml microcentrifuge tube.
- Centrifuge the cells at maximum speed for 15 seconds.
- Resuspend the cells in 200 µl of pre-warmed pulse medium (Dulbecco Modified Eagle Medium DMEM supplemented with 0.20 mM L-cysteine and 0.2 mCi/ml [35S]-methionine). Then, incubate the cells in a water bath at 37oC for 2 minutes.
- Centrifuge for 15 seconds at 4oC at maximum speed.
- Resuspend the pellet in 1.05 ml pre-warmed chase medium (DMEM media supplemented with 7.5% fetal bovine serum and 67 mM L-methionine).
- Incubate the cells in a water bath at 37oC for 20 minutes.
- Take 190 µl aliquots each at 0, 5, 10, 15, 20 minutes.
- Immediately transfer them to 1.5 ml centrifuge tubes containing 1 ml ice-cold DPBS.
- Centrifuge for 15 seconds at 4 o
- Resuspend the pellet in 1 ml non-denaturing lysis buffer (ice-cold)
- Incubate the cell lysate at 4oC for 30 minutes with slow rotation.
- Centrifuge for 15 minutes at 4oC at maximum speed.
- Discard the pellet and keep the supernatant.
Immunoprecipitation
- Wash 30 µl of protein A sepharose twice with 0.5 ml ice-cold Dulbecco’s phosphate-buffered saline DPBS and centrifuge for 1 minute at 3000 x g at 4o
- Resuspend resin in 0.5 ml ice-cold DPBS supplemented with 0.001% Triton X-100 and anti-HA antibody of interest.
- Incubate for 2 hours at 4oC under slow rotation.
- Wash the resin with 0.5 ml ice-cold non-denaturing lysis buffer (0.5% Triton X-100, 50 mM Tris-HCl (pH 7.5), 300 mM NaCl, 5 mM EDTA, and Complete mini, EDTA-free protease inhibitor cocktail) for two times.
- Add 10 µl bovine serum albumin.
- Add cleared cell lysate from each aliquot.
- Incubate for 2 hours at 4oC under slow rotation.
- Wash the resin twice with 0.5 ml ice-cold non-denaturing lysis buffer supplemented with 0.1% instead of 0.5% Triton X-100.
- Wash the resin with 0.5 ml ice-cold DPBS.
- Resuspend the resin in 20 µl of 4x LDS buffer.
- Boil the samples for 5 minutes.
SDS-PAGE and fluorography
- Load 15 µl of each sample on protein mini-gels.
- Run the gel for 3 hours at a constant 50 mA/gel.
- Fix the gels with 10 ml fixation solution (50% methanol and 10% acetic acid) for 30 minutes at room temperature under slow rocking.
- Dry gels in gel drier for 1.5 hours at 80o
- Expose the films to the radioactive gels at room temperature overnight.
Single-cell pulse-chase for nuclear components (Drozdz. & Vaux., 2016)
Cell culture
- Culture the cells in cell culture medium containing Dulbecco’s modified Eagle’s medium, 10 % fetal calf serum, nonessential amino acids, and penicillin and streptomycin (100 U/mL).
- Grow the cells in an incubator at 37 °C with 5 % CO 2 supply and split them on 80 % confluence.
- Plate the cells on specific media and perform transfections according to the standard protocols.
Nano SIMS: preparation of stearic acid D35 stock solution
- Add 2 ml ethanol in 3.2 mg of stearic acid and dissolve.
- Mix it with 400 μL of 100 mM solution of NaOH in ethanol.
- Evaporate alcohol to get fatty acid soaps.
- Dissolve the fatty acid soaps in ultrapure water (0.5 ml) prewarmed to 55 °C and keep in a 55 °C water bath for 10 minutes.
- Dissolve 1 g of bovine serum albumin (fatty acid-free) in 9.5-mL Dulbecco’s Modified Eagle’s Medium (DMEM) and warm up to 55 °C.
- Add the dissolved fatty acid soaps to the bovine serum albumin solution (fatty acid-free) and then vortex for 10 seconds, and subsequently incubated for 10-minutes at 55 °C.
- Filter the stock solution under sterile conditions, aliquot, and keep at -20 °C to make a 1 mM stearic acid D35 stock solution.
Stearic Acid D35 cell labeling and embedding
- Plate mouse preadipocytes on 13-mm plastic coverslips in a 24-well plate at a 50 % confluence rate and leave them overnight.
Note: Coverslips pre-coated with poly-l-lysine, collagen, or more elaborate extracellular matrix preparations can be used depending on the cell type.
- Treat the cells with 10 μM stearic acid D35. Add 100 μL of 1 mM stearic acid stock solution to the 10-mL culture medium (pre-warmed) and vortex. Use it to replace the medium in the 24-well plate (500 μL per well). Leave for 6 hours in the incubator.
- Aspirate the medium and wash the coverslips with 1 mL phosphate buffered saline (PBS). Discard the PBS.
- Fix the cells by adding 500 μL of fixative warmed to 37 °C. Use 4% paraformaldehyde and 1 % glutaraldehyde solution in 100 mM PIPES pH 7.4. Leave for 20 minutes at room temperature.
- Replace the fixative with 500 μL of 2.5 % glutaraldehyde dissolved in 100 mM PIPES pH 7.4.
- Incubate for 1 hour at room temperature and transfer to 4 °C. Leave it overnight.
- Wash the samples thrice with 100 mM PIPES pH 7.4 for 10 minutes.
- Osmicate the cells with 1 % osmium tetroxide in 100 mM PIPES pH 7.4 for 1 hour.
- Wash the samples with deionized water for 20 minutes.
- Incubate the samples in a graded ethanol series, starting from 50 % ethanol for 15 minutes, then 70 % ethanol overnight at 4 °C, then 90 % ethanol for 15 minutes, then 95 % ethanol for 15 minutes, and 100 % ethanol for 2 hours with three solution changes during this time.
- Gradually infiltrate the samples with agar 100 epoxy resin by serial replacement of the resin solution in the coverslip-containing wells. Start with 25 % resin for 1 hour, then 50 % resin for 2 hours, then 75 % resin for 1 hour, and 100 % resin overnight.
- Transfer the samples to fresh 100 % resin for 3 hours and repeat this step once more.
- Resin-embed the layer of cells by inverting the coverslip onto an embedding capsule filled with fresh 100 % resin. Leave for 24 hours at 60 °C to allow polymerization.
- Submerge the blocks in liquid nitrogen and take off the coverslip and leave the cells embedded in the resin.
- Cut the resin block using a razor blade and glass knife to generate a trapezoid end containing the specimen.
- Cut the specimen into semi-thin sections of 0.5 μm using an ultramicrotome with a diamond knife, and mount them on 15-nm platinum-coated coverslips. Place them on a 60 °C heating block and allow to dry for a few minutes.
Backscattered electron imaging and NanoSIMS
- Record the points of interest by optical microscopy before starting backscattered electron (BSE) imaging.
- Transfer the sections to a scanning electron microscope, and acquire BSE images with a 2-kV incident beam with standard aperture (30 μm) and 5-mm working distance.
- Coat the sections with 5 nm of platinum in a high-resolution sputter coater to make the surface conductive for NanoSIMS imaging.
- A small region of the sample is revealed by uncoating to remove the material from the section for 2 H − and 1 H − mass analysis using the cesium ion beam.
- Use small apertures (D1 = 3 or D1 = 4) for single cells imaging to match the primary beam size with the pixel size. Tune the instrument for 2 H − and 1 H – for morphological information and calculate the 2 H/ 1 H ratio. Collect and process NanoSIMS images with a dwell time of 30,000 μs per pixel for a 256 × 256-pixel image.
- Manually align the BSE and NanoSIMS images for the identification of regions containing structures of biological interest.
Applications
Analysis of the in vivo assembly of bacteriophage T4 tail (Ferguson. & Coombs., 2000)
The pulse-chase analysis is a non-invasive tool to determine the assembly pathway of the complex tail of bacteriophage T4 virus. In the study, bacteriophage T4 mutants defective in the head assembly were used to infect the E. coli cultures to study tail assembly in isolation. The bacterial cultures were labeled with [3H] leucine on the onset of late protein synthesis. After the complete tail began to accumulate at a constant rate, the bacterial cultures were pulsed with [35S] methionine, and then chased. Completed tails were obtained and purified at one-minute intervals for the next 30 minutes. It was found that the closer the assembly point to the end of the pathway, the sooner the chase appears, presenting the assembly cascade. The results showed that the tail completion proteins such as gp18 (tail sheath) and 19 (tail tube) show the earliest inflection as compared to other tail proteins. The study validated the application of pulse-chase analysis to analyze and assess the assembly of viral components.
Assessment of protein maturation and degradation (Jansens. & Braakman., 2003)
The pulse-chase analysis is a powerful technique to analyze protein maturation and degradation. Using the short pulses, the whole process of protein synthesis to degradation can be determined in a natural environment. The technique has been widely used for endogenous and viral proteins including thyroglobulin, vesicular stomatitis virus, influenza hemagglutinin, envelope proteins, and immunoglobulins. This biochemical approach is a cutting-edge analysis method to study a variety of processes including protein folding, post-translational modifications, endogenous and intracellular transport, and the rate of protein degradation.
Assessment of MHC class II biosynthesis, maturation, and peptide loading (Hou. et al. 2013)
The pulse-chase analysis is a widely used technique for the exploration of the synthesis, processing, and transport of proteins. In the research, the pulse-chase experiment was used to study the major histocompatibility complex (MHC) class II synthesis, maturation, trafficking, and peptide loading in human Epstein-Barr virus-transformed B-lymphoblastoid cell lines (B-LCL). The MHC class II glycoproteins bind heterogeneous mixtures of peptides presented on the surface of antigen-presenting cells for the inspection of CD4+ T helper lymphocytes. The results obtained from these experiments can illustrate information to track changes in the molecular associations, an abundance of radiolabeled proteins, and the conformation-sensitive monoclonal antibodies (mAbs).
Biosynthesis, targeting, and processing of lysosomal proteins (Pohl. & Hasilik., 2015)
The radioactive amino acids and modifier groups are incorporated into proteins to study their life cycle and various modifications. Lysosomal enzymes can also be detected and characterized using pulse-chase analysis. The pulse-chase labeling provides a detailed insight into organelle-specific modifications of lysosomal proteins. For this, an antibody against lysosomal protease is used as a reference. In the study, cathepsin Z was synthesized as a larger proenzyme containing two N-linked oligosaccharides which mature to a shorter single-chain enzyme with oligosaccharides. The pulse-chase experiment demonstrates the conversion of the precursor into the mature form. The technique could also be used to study the deglycosylation of metabolically labeled proteins and the alterations in the apparent size of the attached glycopeptides.
Precautions
- Prepare fresh pulse-chase media before use.
- During lysis, it is essential to maintain the nucleus intact.
- If the protein expression is relatively low, then the cells can be transfected with a plasmid encoding the protein behind the promotor or a lipid mixture.
- Separate the folding process from translation if the effects of certain conditions such as ATP depletion are tested.
- Cycloheximide should be added in chase medium if the kinetics of the process has to be studied.
Strengths and Limitations
- The pulse-chase analysis is a powerful technique used for the assessment of protein synthesis, trafficking, and degradation.
- The technique could also be used to monitor the binding of proteins and the kinetics of the processes.
- The pulse-chase method is widely used to monitor the assembly cascade of viruses.
- The method should be readily applicable to the detection and assessment of multicomponent regulatory complexes and macromolecular machines accessible to the biochemical and biophysical investigation.
- The approach is limited to monitoring the kinetics of stable interactions.
- The labeling of the proteins could be challenging for low-abundant proteins.
References
- A. Jansens., & Braakman., I. (2003). Pulse-chase labeling techniques for the analysis of protein maturation and degradation. Methods Mol Biol, 232, 133-45.
- P. Ferguson., & Coombs., H. D. (2000). Pulse-chase analysis of the in vivo assembly of the bacteriophage T4 tail. J Mol Biol, 297(1), 99-117.
- Drozdz., & Vaux., J. D. (2016). Methods for Single-Cell Pulse-Chase Analysis of Nuclear Components. Methods Mol Biol, 1411, 159-76.
- Magadán, J. G. (2017). Radioactive Pulse-Chase Analysis and Immunoprecipitation. Bio Protoc, 4(8).
- Pohl., & Hasilik., A. (2015). Biosynthesis, targeting, and processing of lysosomal proteins: pulse-chase labeling and immune precipitation. Methods Cell Biol, 126, 63-83.
- Hou., C. H. Rinderknecht., A. V. Hadjinicolaou., R. Busch., & Mellins, E. (2013). Pulse-chase analysis for studies of MHC class II biosynthesis, maturation, and peptide loading. Methods Mol Biol, 960, 411-432.